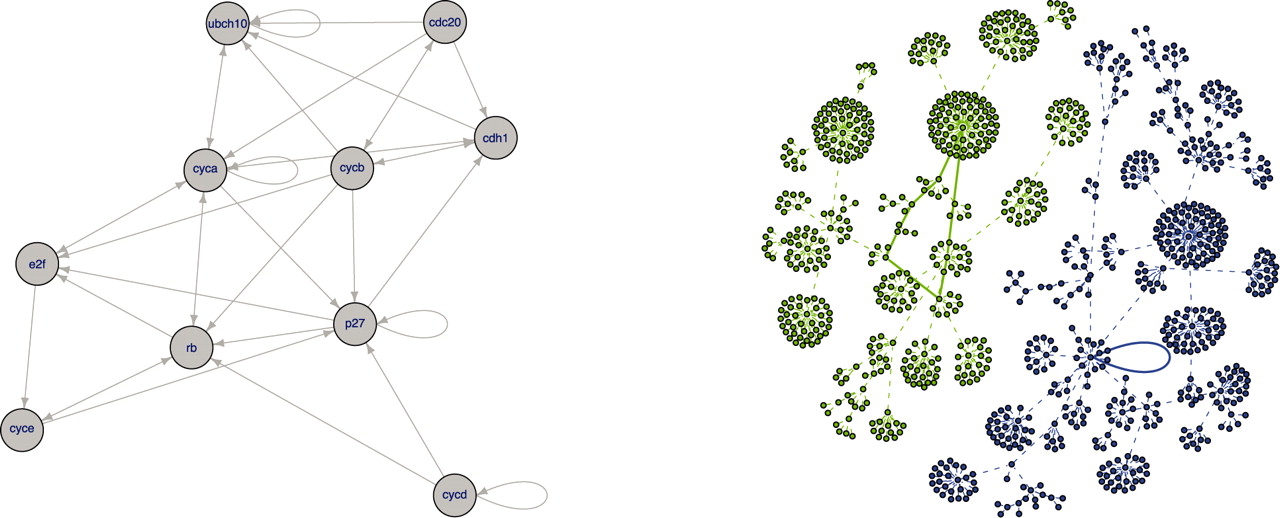
BNToolkit is a C++ tool developed at Politecnico di Torino able to simulate and analyze Boolean Networks describing gene regulatory activities. One of the most important characteristics of the tool is its flexibility that anables to preciselly model important regulatory activities such as post-transcriptional regulation carried out by miRNAs.
BNToolkit is able to analyze a network computing the network attractors and the network state space and producing an output in a standard xgmml format.
Developers

How to cite BNToolkit
The SysBio development team has invested a lot of time and effort in creating BNToolkit as it is today. Please give credit where credit is due and cite BNToolkit when you use it for your research activities.
BNToolkit can be cited through this papers:
Benso A., Di Carlo S., Rehman H.U. ,Politano G.,Savino A., Squillero G.,Vasciaveo A., Benedettini S. (2013)
Accounting for Post-Transcriptional Regulation in Boolean Networks Based Regulatory Models. In: International Work-Conference on Bioinformatics and Biomedical Engineering, IWBBIO 2013, Granada, ES, 18-20 March , 2013. pp. 397-404
BibTEX
@article{benso2013accounting,
title={Accounting for post-transcriptional regulation in boolean networks based regulatory models},
author={Benso, Alfredo and Di Carlo, Stefano and Rehman, Hafeez Ur and Politano, Gianfranco Michele Maria and Savino, Alessandro and Squillero, Giovanni and Vasciaveo, Alessandro and Benedettini, Stefano},
year={2013},
publisher={Copicentro Editorial}
}
Benso A.; Di Carlo S.; Politano G.; Savino A.; Vasciaveo A. (2014)
An Extended Gene Protein/Products Boolean Network Model Including Post-Transcriptional Regulation. In: THEORETICAL BIOLOGY AND MEDICAL MODELLING, vol. 11(Suppl 1) n. S5, pp. 1-17. – ISSN 1742-4682 DOI: 10.1186/1742-4682-11-S1-S5
BibTEX
@Article{Benso2014,
author="Benso, Alfredo
and Carlo, Stefano Di
and Politano, Gianfranco
and Savino, Alessandro
and Vasciaveo, Alessandro",
title="An extended gene protein/products boolean network model including post-transcriptional regulation",
journal="Theoretical Biology and Medical Modelling",
year="2014",
volume="11",
number="1",
pages="1--17",
issn="1742-4682",
doi="10.1186/1742-4682-11-S1-S5",
url="http://dx.doi.org/10.1186/1742-4682-11-S1-S5"
}
Gianfranco Politano, Alessandro Savino, Alfredo Benso, Stefano Di Carlo, Hafeez Ur Rehman, Alessandro Vasciaveo, Using Boolean networks to model post-transcriptional regulation in gene regulatory networks, Journal of Computational Science, Volume 5, Issue 3, May 2014, Pages 332-344, ISSN 1877-7503, http://dx.doi.org/10.1016/j.jocs.2013.10.005.
(http://www.sciencedirect.com/science/article/pii/S1877750313001129)
BibTEX
@article{Politano2014332,
title = "Using Boolean networks to model post-transcriptional regulation in gene regulatory networks ",
journal = "Journal of Computational Science ",
volume = "5",
number = "3",
pages = "332 - 344",
year = "2014",
note = "",
issn = "1877-7503",
doi = "http://dx.doi.org/10.1016/j.jocs.2013.10.005",
url = "http://www.sciencedirect.com/science/article/pii/S1877750313001129",
author = "Gianfranco Politano and Alessandro Savino and Alfredo Benso and Stefano Di Carlo and Hafeez Ur Rehman and Alessandro Vasciaveo",
}
Downloads
Download the BNToolkit. Build instructions are provided in the README file.
Download the Cytoscape plugin required to visualize the xgmma output.